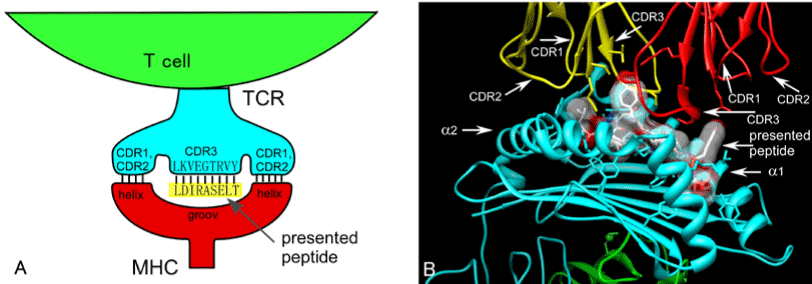
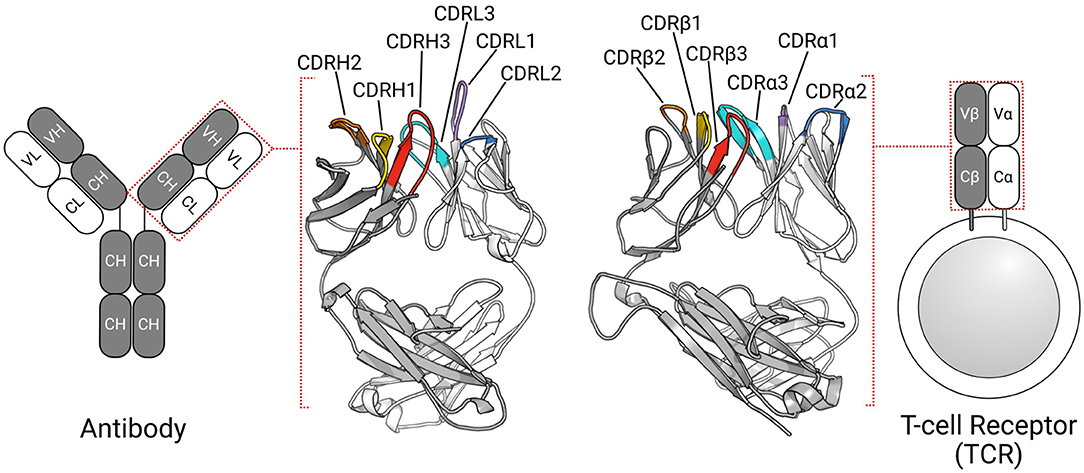
Antigen recognition in adaptive immunity Mechanism of diversity of antigen receptors Maturation and selection of B lymphocytes Maturation and selection. - ppt download
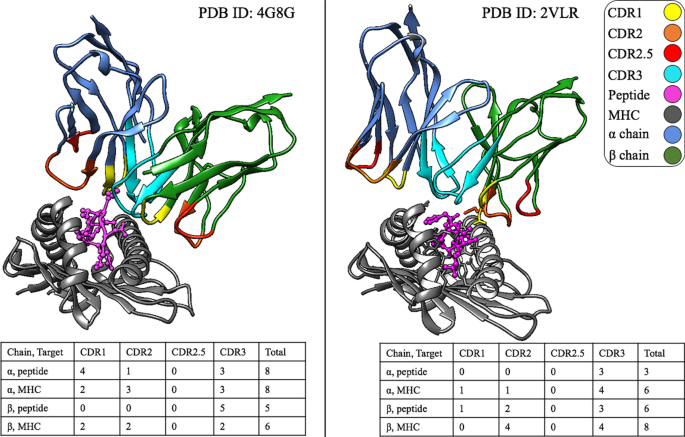
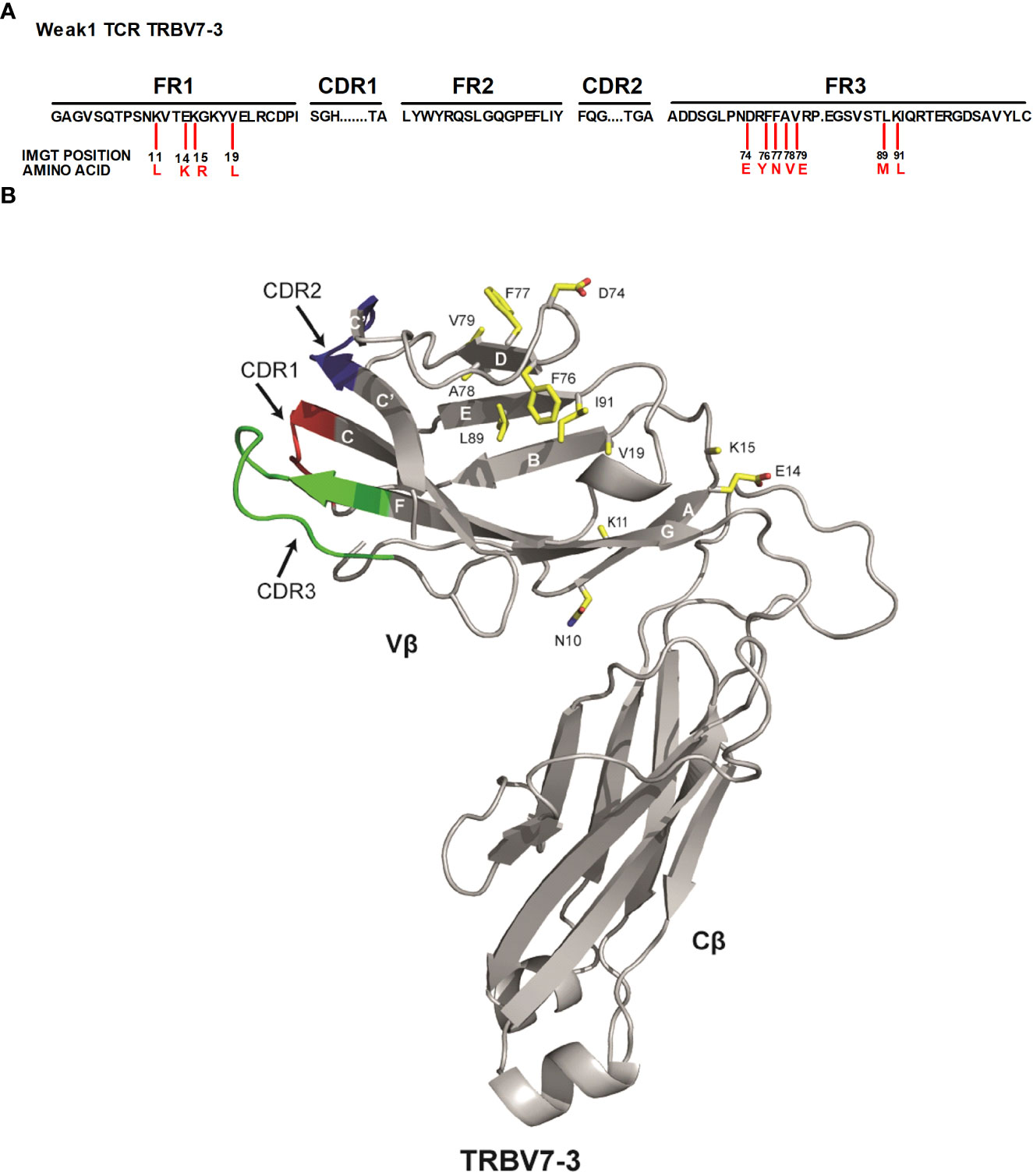
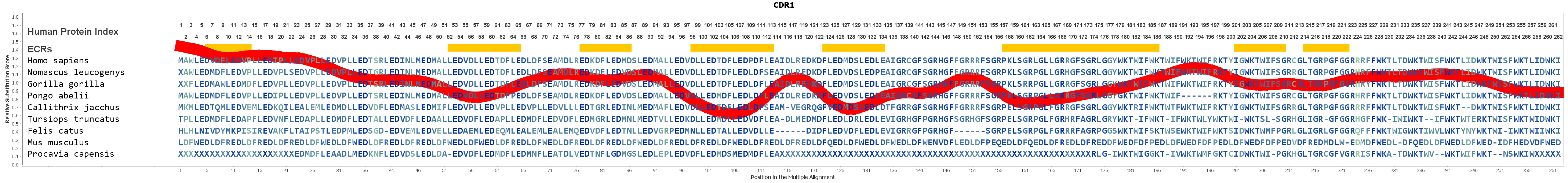
SwarmTCR: a computational approach to predict the specificity of T cell receptors | BMC Bioinformatics | Full Text
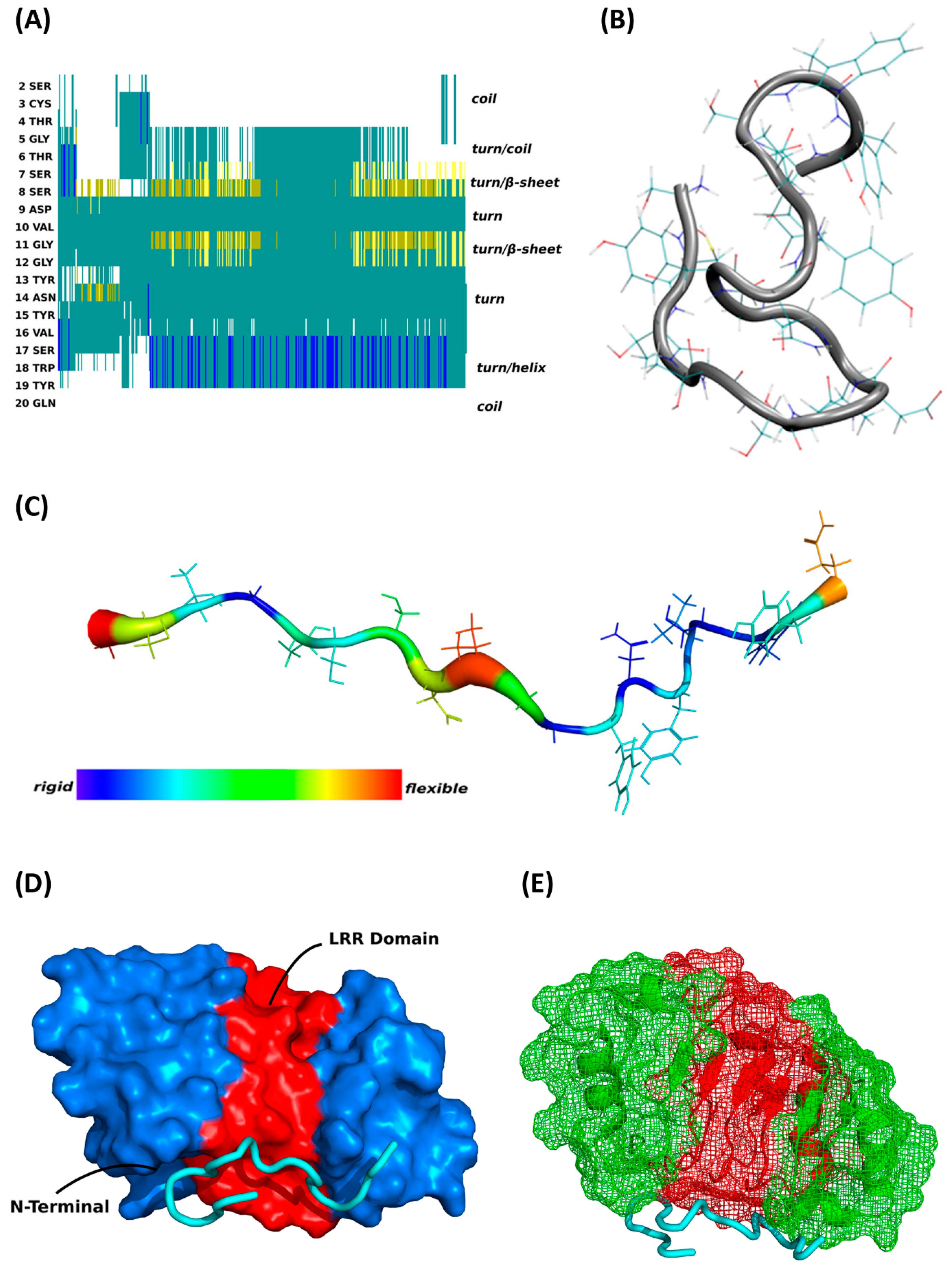
Biomolecules | Free Full-Text | Glaucoma-Associated CDR1 Peptide Promotes RGC Survival in Retinal Explants through Molecular Interaction with Acidic Leucine Rich Nuclear Phosphoprotein 32A (ANP32A)
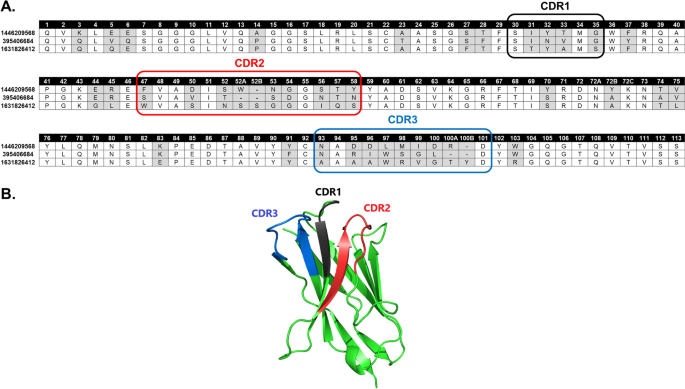
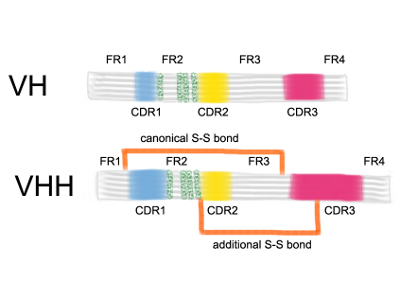
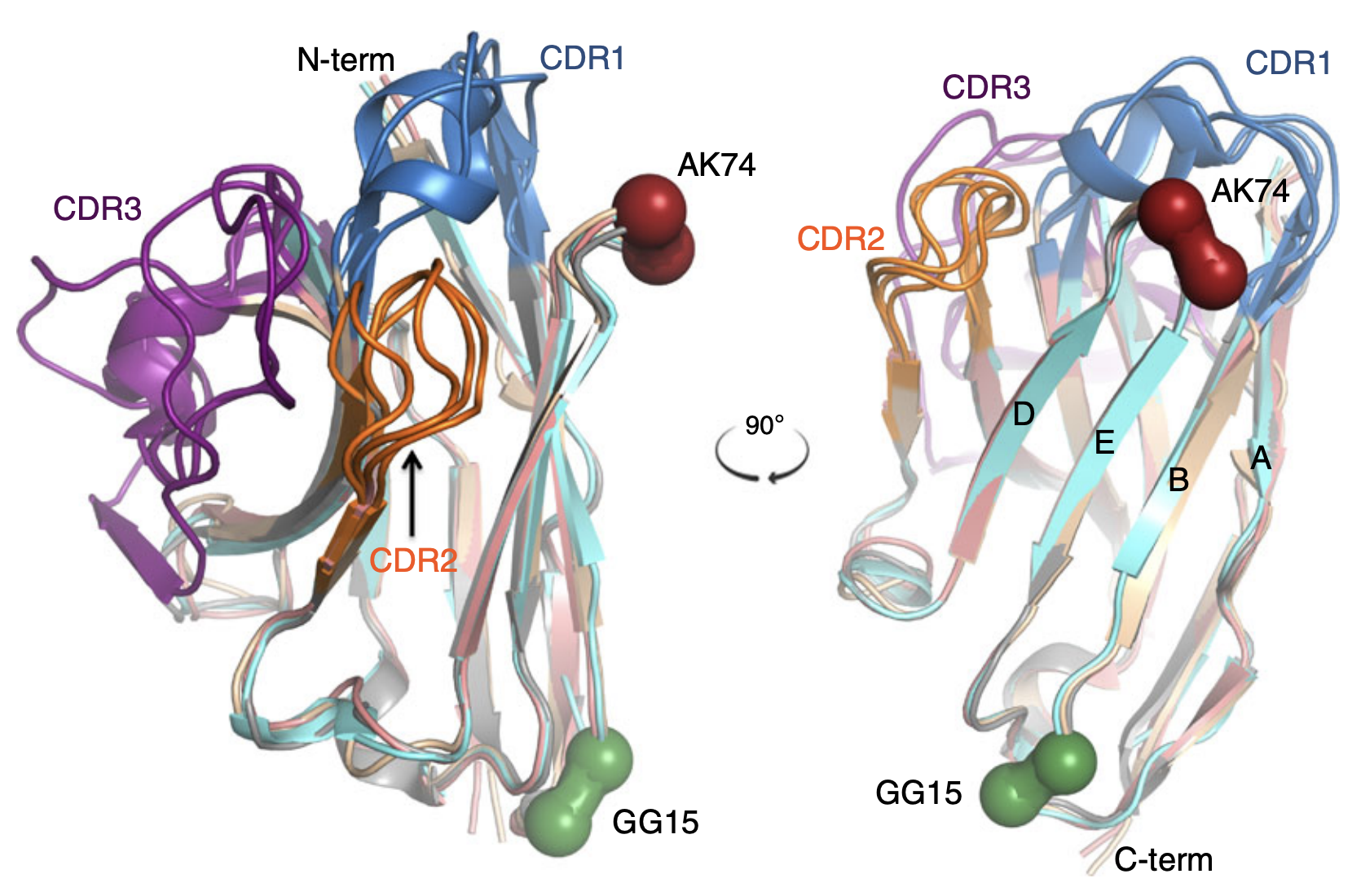
Studying the characteristics of nanobody CDR regions based on sequence analysis in combination with 3D structures | Journal of Genetic Engineering and Biotechnology | Full Text
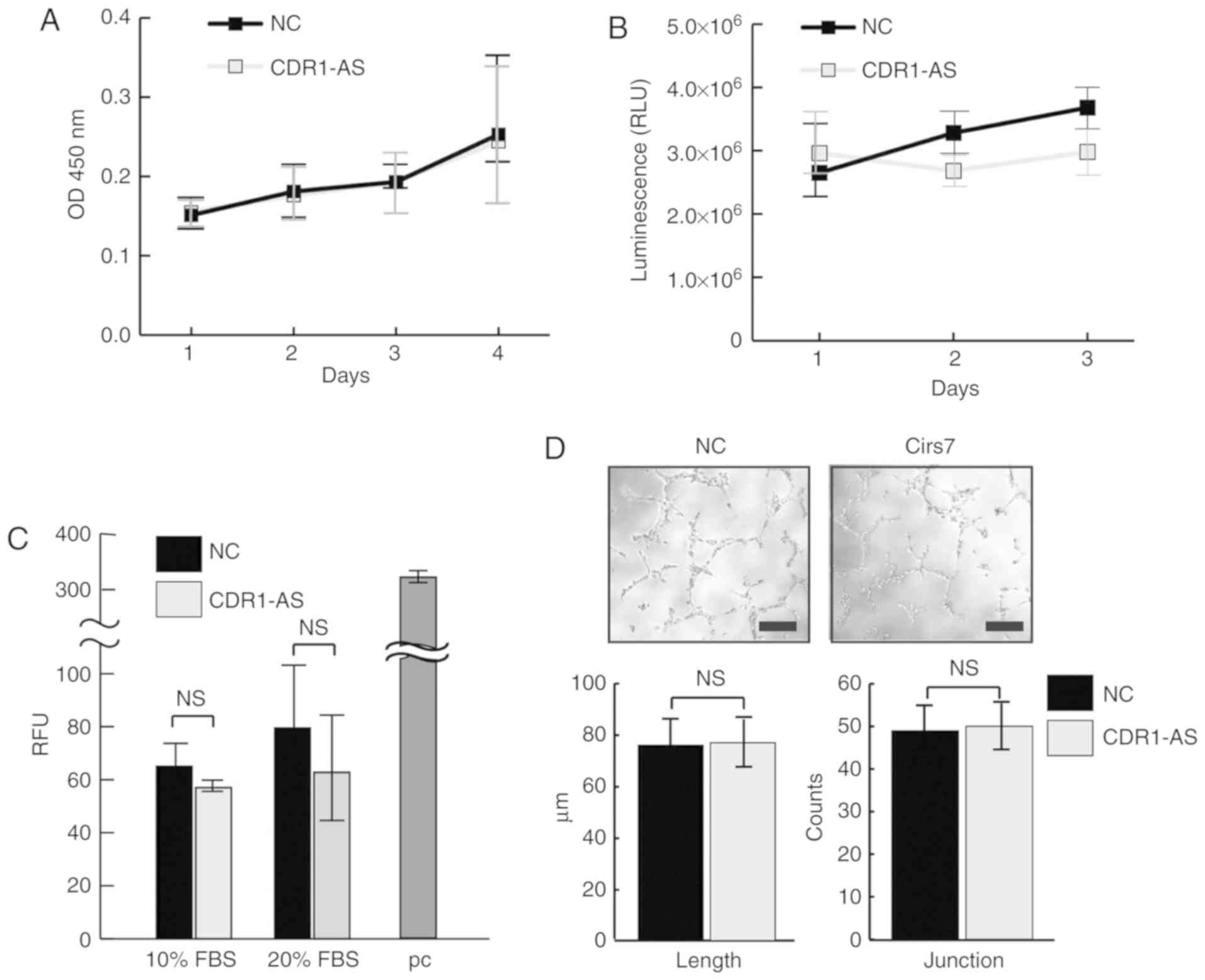
Expression of circular RNA CDR1‑AS in colon cancer cells increases cell surface PD‑L1 protein levels
Uncovering the Principles of Light-Switchable Protein Design through Genome-Wide Interrogation of Light Oxygen Voltage Domain Insertion Sites in Yeast | Institute for Collaborative Biotechnology (ICB) | UCSB, MIT and Caltech.

Diversi®cation of VH CDR1, CDR2 and CDR3 to make intrabody libraries. (... | Download Scientific Diagram

Hiding in plain sight: structure and sequence analysis reveals the importance of the antibody DE loop for antibody-antigen binding | bioRxiv

ModiBodies: A computational method for modifying nanobodies to improve their antigen binding affinity and specificity | bioRxiv